2. Protein Switches
One-pot Colorimetric Detection of Molecules Based on Proximity Proteolysis Reaction
Biosensors and Bioelectronics, 2021, 188, 113349
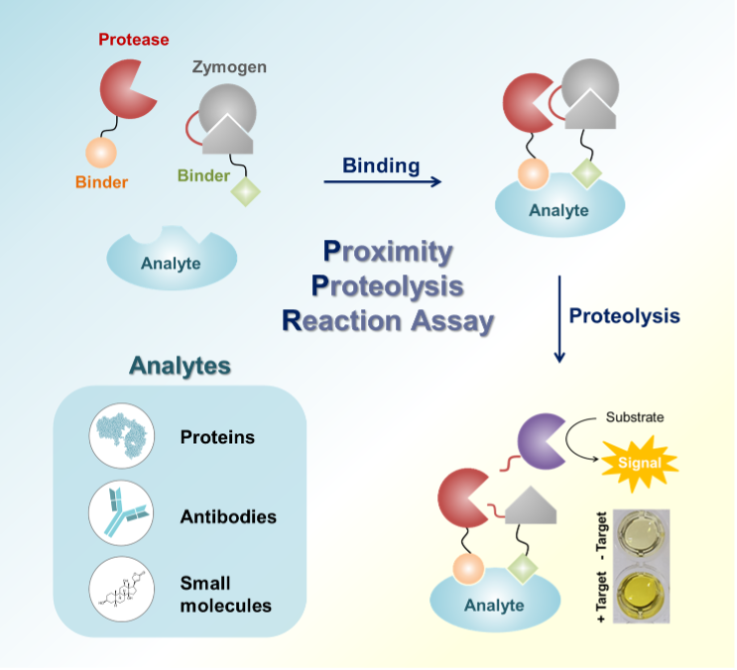
Various types of molecules serve as biomarkers of diseases, and numerous methods have been reported to detect and quantify them. Recently, research efforts have been made to develop point-of-care (POC) tests, which contribute to early diagnoses of diseases, particularly in resource-limited settings. An assay performed in a homogeneous phase is an obvious route to develop these methods. Here, simple homogeneous methods based on proximity proteolysis reactions (PPR) are reported to detect biological molecules. A typical PPR system has been designed such that the proteolysis reaction between protease and zymogen is enhanced in the presence of a target analyte. The activated zymogen generates a color signal by hydrolyzing a chromophore. A protease and zymogen are linked to target binders using specific hybridization between complementary single-stranded DNAs, and several molecules, including proteins, antibodies, aptamers, and small molecules, are used as target binders. The developed assay methods successfully detected several kinds of analytes at subnanomolar concentrations with the one-step procedure and color signal. The modular design of the PPR-based assay will enable the development of simple POC diagnostics for various biomarkers
Nucleic Acid Detection by a Target-Assisted Proximity Proteolysis Reaction
ACS Sensors, 2018, 3, 2066-2070
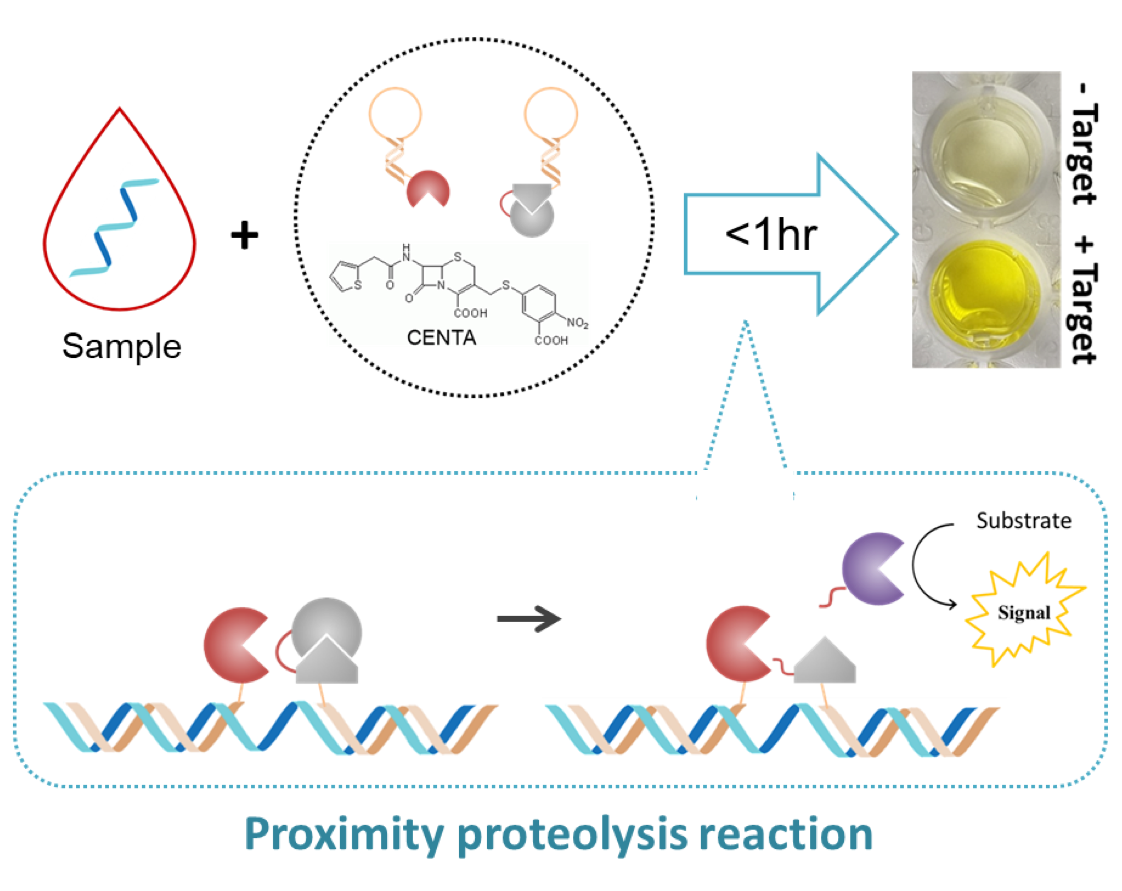
Nucleic acid analysis plays an important role indiagnosing diseases as well as understanding biology. Despite advancesin technology, there is still a need to develop a rapid and simplemethod to detect specific nucleic acids, especially in remote locationsand low-resource cases. Here, we proposed a proximity proteolysisreaction in which the reaction between protease and zymogen isenhanced in the presence of a target molecule. The pair of proteins wassite-specifically modified with oligonucleotides, and the conjugateswere used to develop a method of detecting nucleic acids. Target DNAand RNA could be detected in less than 1 h at sub-nanomolarconcentrations based on an absorbance signal. The assay method wasresistant to interference by biological matrixes, and its sensitivity couldbe improved when combined with an isothermal nucleic acidamplification method. The results demonstrated the feasibility of this proximity proteolysis reaction as a new platformtechnology for detecting specific nucleic acid sequences.
Engineering beta-lactamase zymogens for use inprotease activity assays
ChemComm, 2014, 50, 10155-10157
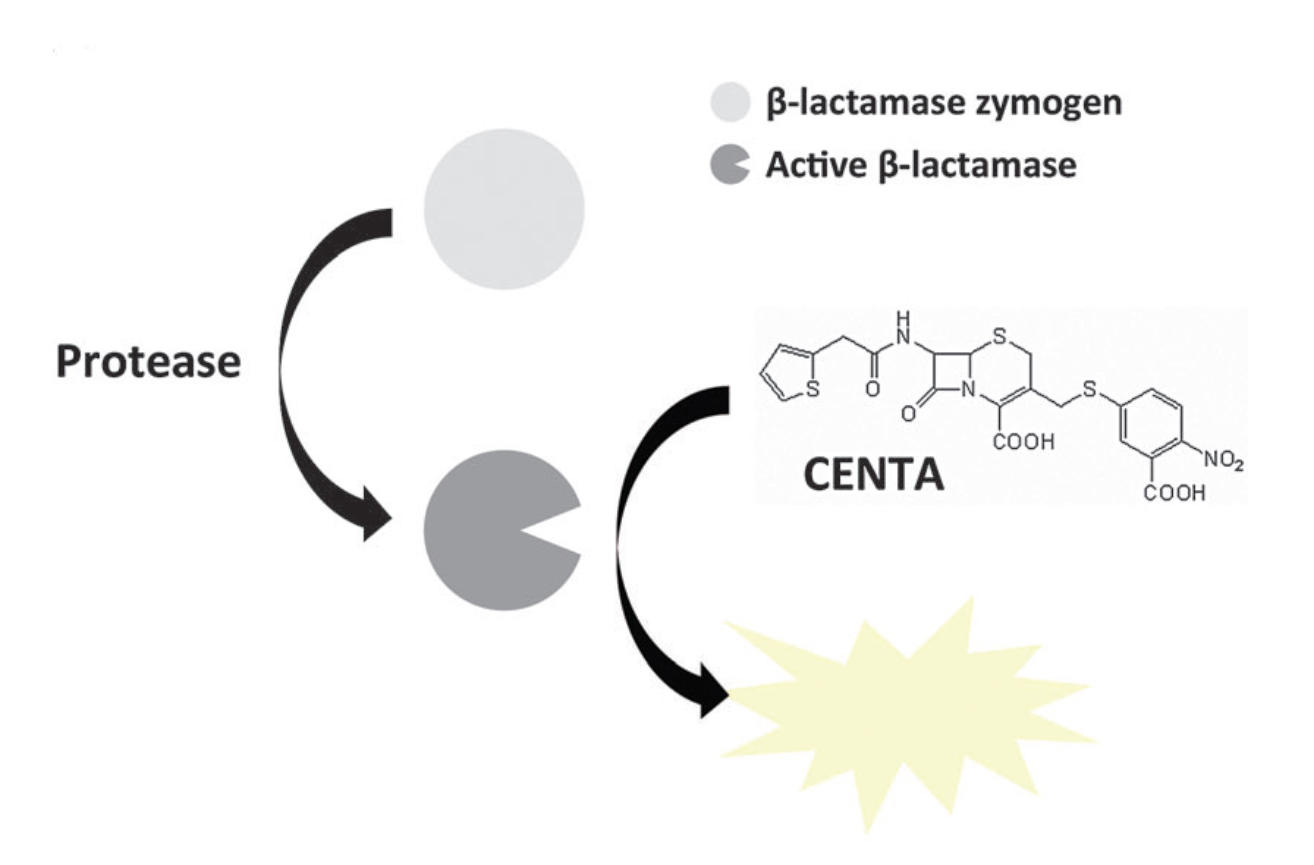
In this study, the concept of autoinhibition, a mechanism of proteinactivity regulation, was applied to the design and engineering of ab-lactamase zymogen. Using this zymogen, a sensitive proteaseassay method was developed in which activation of the zymogen byproteases produces an amplified absorbance signal. The approachreported here can be adapted for engineering of zymogens asbiological sensors and components of synthetic signaling pathways.

Synthetic Protein Engineering Laboratory
Department of Molecular Science and Technology, Ajou University
206 Worldcup-ro, Yeongtong-gu, Suwon, 16499 Korea